How to Measure Contingency-Coefficient (Association Strength)
How to Measure Contingency-Coefficient, when the hypothesis of independence of attributes in a contingency table is rejected by performing a chi-square test, ensures the association between two attributes.
Such kinds of situations are interested to calculate the strength of association and it is a desideratum. For this, a measure is known as the coefficient of contingency.
The measure of contingency was developed by Karl Pearson in 1904.
The coefficient of contingency is denoted as C.
C=sqrt(χ2/n+ χ2)
Where n is the sample size. The value of C lies between 0 and 1 and never attains 1.
When C=0 indicates complete dissociation. A value near 1 indicates a high degree of association.
In most cases, C calculated when the null hypothesis rejected.
Chi square distribution examples
Introduction to Chi-Square Test
The Chi-Square test in R is a statistical method used to determine if two categorical variables have a significant correlation between them.
The two variables are selected from the same population and labeled as categorically.
Syntax: chisq.test() is a function used to perform the test in R
We reject the null hypothesis if the p-value that comes out in the result is less than a significance level, which is 0.05 usually.
H0: The two variables are independent.
H1: The two variables relate to each other.
Getting Data
data("mtcars")
table(mtcars$carb, mtcars$cyl)Contigency table from mtcars
4 6 8 1 5 2 0 2 6 0 4 3 0 0 3 4 0 4 6 6 0 1 0 8 0 0 1
Visulization
library(gplots) dt<-table(mtcars$carb, mtcars$cyl) balloonplot(t(dt), main ="Balloon Plot", xlab ="", ylab="", label = FALSE, show.margins = FALSE)

Let’s calculate the chi-square value based on chisq.test function
chisq.test(mtcars$carb, mtcars$cyl)
Pearson’s Chi-squared test
data: mtcars$carb and mtcars$cyl X-squared = 24.389, df = 10, p-value = 0.006632 Warning message: In chisq.test(mtcars$carb, mtcars$cyl) : Chi-squared approximation may be incorrect
Now you can see the warning because some of the cell frequencies are less than 5.
According Chi-Square test significant difference was observed between tested attributes. However, will check fisher’s exact results also.
Fisher’s Exact Test for Count Data
fisher.test(mtcars$carb, mtcars$cyl) data: mtcars$carb and mtcars$cyl p-value = 0.0003345 alternative hypothesis: two.sided
There is no changes in the inferences significant difference was observed between variables, now will check the association of variables.
Let’s visualize Pearson residuals using the package corrplot:
library(corrplot) chisq<-chisq.test(mtcars$carb, mtcars$cyl) corrplot(chisq$residuals, is.cor = FALSE)
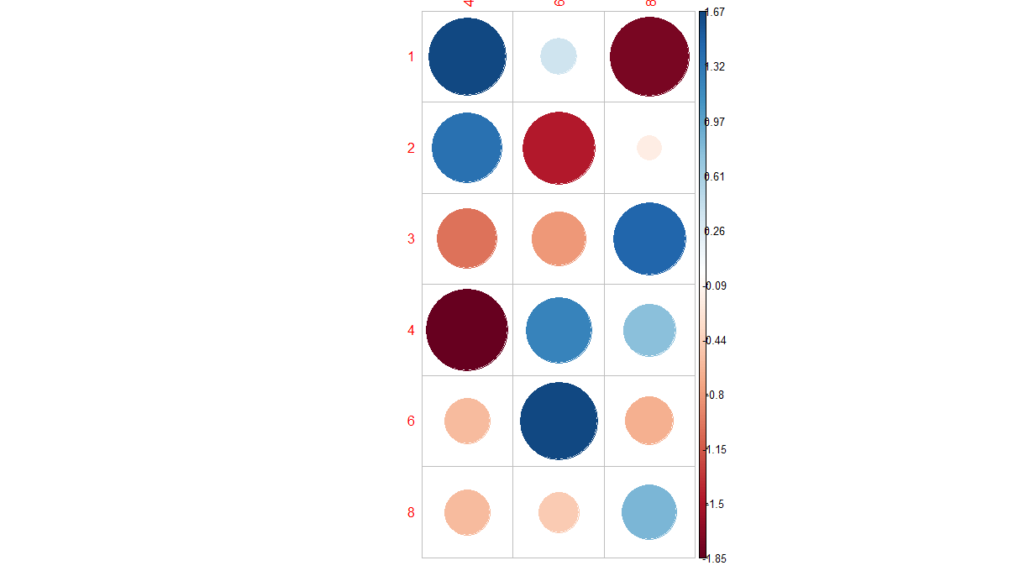
For a given cell, the size of the circle is proportional to the amount of the cell contribution.
The sign of the standardized residuals is also very important to interpret the association between rows and columns.
Positive residuals are in blue and Negative residuals are in red. Based on the above plot, association observed as high.
Customer Segmentation K Means Cluster
Contingency Coefficient
Assocs function fron DescTools returns different association measures simultaneously.
library(DescTools) Assocs(dt)
estimate lwr.ci upr.ci Phi Coeff. 0.8730 - - Contingency Coeff. 0.6577 - - Cramer V 0.6173 0.1778 0.7592 Goodman Kruskal Gamma 0.6089 0.3745 0.8432 Kendall Tau-b 0.4654 0.2846 0.6463 Stuart Tau-c 0.4834 0.2981 0.6687 Somers D C|R 0.4319 0.2650 0.5989 Somers D R|C 0.5015 0.2935 0.7095 Pearson Correlation 0.5746 0.2825 0.7692 Spearman Correlation 0.5801 0.2900 0.7725 Lambda C|R 0.4444 0.1198 0.7691 Lambda R|C 0.2727 0.0000 0.5570 Lambda sym 0.3500 0.1128 0.5872 Uncertainty Coeff. C|R 0.4782 0.3693 0.5870 Uncertainty Coeff. R|C 0.3387 0.2752 0.4022 Uncertainty Coeff. sym 0.3965 0.3220 0.4710 Mutual Information 0.7353 - -
From the above table Contingency Coefficient value observed as 0.6577, indicates a high association between the tested variables.
Summary
Based on the above analysis, a significant difference was observed between the variables (ie variables are related) and strength of association between the variable is high.